Alfonso Valencia has led this project to discover the location of changes and their relation with cellular diversity and cancer origin
A team of scientists from different institutions led by the BSC researcher Alfonso Valencia and by the Newcastle University researcher Daniel Rico has developed a method to detect the epigenome areas where the changes that give rise to cellular diversity originate. These changes are also related to the cancer origin and development. In the study, which has been published in Nucleic Acids Research, they have developed a computational method that has been applied in hematopoiesis, the process of blood formation from stem cells.
The system allows the integration of a variety of epigenomic datasets to classify different samples and automatically identify genomic regions in which epigenomic changes affect the definition of cell type. Furthermore, they showed how epigenomic changes in these regions can be related to different types of leukemias.
The human genome is a dull sequence of letters but it becomes alive thanks to the help of the epigenome. The genome is like a book in which letters follow each other without empty spaces, full stops or commas. It would be very hard to read this book, utterly impossible to understand it. However, with the simple addition of punctuation marks (commas, colons, full stops) we can read and understand the meaning of that apparently dull sequence of letters. This is the great task accomplished by the epigenome, which is composed of chemical changes on the DNA that allow us and the cell to understand how to read and interpret the genome.
For this reason, studying the epigenome is paramount to understanding how development can give rise to the huge variety of cell types forming tissues and organs, all starting from a single cell and a single genome. The epigenome is also often involved in explaining how a healthy cell gives rise to a tumour after a malignant transformation. However, despite the large collections of data available until now, locating those regions of the genome with chemical changes (punctuation marks) and type of changes remains a challenge for the scientific community.
These chemical changes are produced during development and throughout life as an effect of external factors such as lifestyle and they can be triggers for diseases such as cancer. Hence, identifying epigenetic biomarkers is fundamental to enable new diagnostic and therapeutic strategies.
Molecular classifications have been widely used taking into account gene expression levels as biomarkers, using their on/off state. This method lets researchers identify those regions that are involved in regulating the on/off gene state like biological switches. These regions could be used as epigenetic biomarkers that complement the actual molecular classifications.
“The development of this type of methods is very important because up to now differences between cell types had mostly been characterised at the levels of genes that are either switched on or off, that is the final product of the epigenomic regulation, but we did not know where the switches for these genes were (encoded in the epigenome). This acquired knowledge is fundamental to enable new therapies based on acting on the correct switches in cases where the cell loses control in diseases such as cancer. Understanding this level of regulation will take us one step further in the personalized medicine agenda”, says Enrique Carrillo, co-first author of the study and researcher at CNIO.
On the other hand, the joint effort within member projects of IHEC (International Human Epigenome Consortium) in the last few years has generated an impressive data collection that allows us to delve deeper into the study of cellular regulation and its deregulation. According to Alfonso Valencia, Life Sciences Department Director at BSC: “IHEC has made 9000 epigenomes openly available and combining this with the most recent update to the MareNostrum4 Supercomputer will enable us to design new analysis pipelines and more advanced computational methods that will deepen our knowledge and accelerate our path towards better controlling health. This implies improvements in diagnostic and therapeutic tools for various diseases, as well as defining improved and healthier lifestyles”.
All methods and results produced in this work are publically accessible and will allow the scientific community to identify new epigenetic biomarkers, which may be used in Personalised Medicine diagnosis, and treatment approaches.
These are the institutions that have been involved in this project: Centro Nacional de Investigaciones Oncológicas (CNIO), the Institut de Biologia Evolutiva (IBE: CSIC-UPF), Barcelona Supercomputing Center (BSC), the Institut d'Investigacions Biomèques August Pi i Sunyer (IDIBAPS) and the Institute of Cellular Medicine (Newcastle University). This work was funded by the European Union (FP7/2007–2013, 282510; BLUEPRINT), the Spanish Ministry for Economy and Competitiveness (BFU2015–71241-R) and the philanthropic association “ Friends of CNIO”.
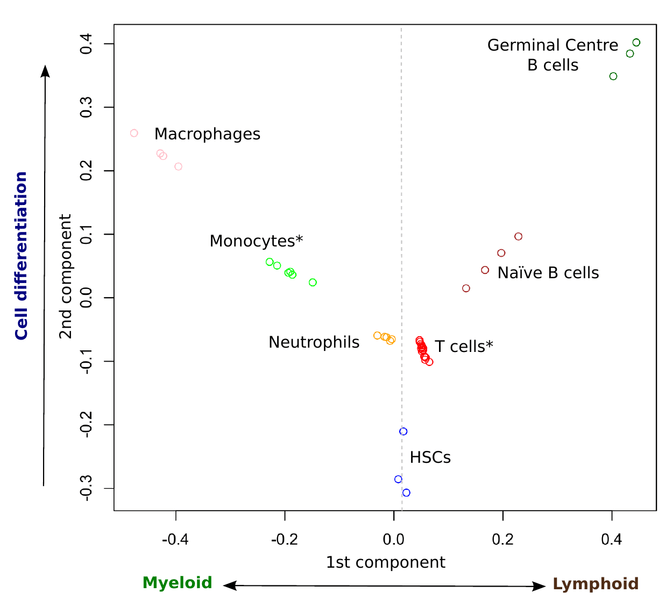
Figure: Distribution of blood cell samples according to their epigenomic profiles produced with the new computational method.
Reference of the study:
Enrique Carrillo-de-Santa-Pau, David Juan, Vera Pancaldi, Felipe Were, Ignacio Martin-Subero, Daniel Rico, Alfonso Valencia, on behalf of The BLUEPRINT Consortium; Automatic identification of informative regions with epigenomic changes associated to hematopoiesis. Nucleic Acids Res 2017 gkx618. doi: 10.1093/nar/gkx618

