The BSC has played a leading role in introducing researchers to collection of tools to explore PCAWG data.
BSC has created one of this tools, PCAWG-Scout, which allows users to run their own analyses on-demand
Nature Communications publishes the research
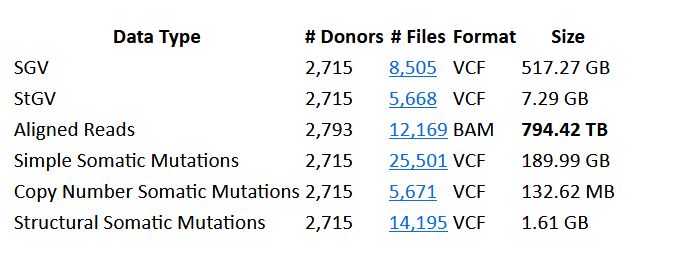
The Pan-Cancer Analysis of Whole Genomes (PCAWG) is an invaluable resource to understand the complex biology of cancer. Its 2,658 samples, chosen to represent 47 different tumor types, were analyzed in depth using whole genome sequencing. The project involved more than 1,300 scientists and clinicians from around to world to produce, process, and interpret this vast data set, nearing 800TB in size.
Now, five institutions involved in the project provide five different data exploration and visualization tools that allow researchers to peer into this complex dataset. In an effort spearheaded by BSC and coordinated also by the UCSC (University of California at Santa Cruz), and with the participation of other prominent institutions like the EMBL- EBI (European Bioinformatics Institute), the OICR (Ontario Institute for Cancer Research) and the Harvard Medical School, they have produced A user guide to the online resources for data exploration, visualization, and discovery for the Pan-Cancer Analysis of Whole Genomes project (PCAWG).
This document, published in Nature Communications, illustrates the different strengths of their corresponding portals, PCAWG-Scout, UCSC Xena, ICGC Data Portal, Expression Atlas, and Chromothripsis Explorer, and demonstrate how they complement each other in understanding more deeply cancer biology.
The paper details use cases and analyses for each tool, shows how they incorporate outside resources from the larger genomics ecosystem, and demonstrates how the tools can be used together to understand the biology of cancers more deeply.
All of the tools aim to streamline analysis and visualization by pre-loading the PCAWG data so that users do not need to locate, curate, or manage the data and by making the tools accessible through a web interface. Each of these five tools also integrates other genomics datasets and tools that provide context and insight for interpretation of patterns in the PCAWG data, helping this resource fully realize its potential.
Miguel Vázquez, Genome informatics group leader at BSC, and senior author of the paper, says that “researchers need tools that show them this data in context, so that interesting patterns are revealed. The interactive nature of these portals offers researchers the opportunity to raise and test hypotheses. What we wanted to highlight in this work was how the complementary ways to immerse yourself in the data that these tools offer reveal interesting phenomena that would otherwise be hidden”.
Five resources to analise the data
The five resources covered in this paper provide a different perspective and focus to the PCAWG data.
- PCAWG-Scout, developed by BSC, allows users to run their own analyses on-demand, including prediction of cancer driver genes, differential gene expression analysis, recurrent structural variations calling, survival analysis, pathway enrichment, visualization of mutations on a protein structure, mutational signatures detection, and recommendations for possible therapies (based on the CNIO’s PanDrugs resource).
- The ICGC Data Portal serves as the main entry point for accessing all PCAWG data and can also be used to explore PCAWG consensus simple somatic mutations, including point mutations and small indels, each by their frequencies, patterns of co-occurrence, mutual exclusivity, and functional associations.
- UCSC Xena integrates diverse types of genomic and phenotypic/clinical information at the sample level across the large number of samples, enabling rapid examination of patterns within and across data types.
- The Chromothripsis Explorer visualizes genome-wide mutational patterns, with a focus on complex genomic events, e.g., chromothripsis and kataegis. This is achieved through interactive Circos plots for each tumor with different tracks that correspond to allele-specific copy number variants, somatic structural variations, simple somatic mutations, indels, and clinical information.
Dealing with 800TB of dataThe management of this huge amount of data is very complex. Moving them is complicated due to their volume and delicate because of how sensitive they are to the privacy of the donors. Romina Royo, researcher in the INB Computational Node 2 group, states: “The analysis that converts the vast volume of raw data into processed data is a complex process that requires enormous computational resources and encounters numerous difficulties. For example, tumors that have undergone catastrophic events in the structure of DNA are a problem for the analysis software, as when they try to untangle the data them they become themselves entangled in loops that can greatly shoot up the computation time”. This first analysis phase required the collaboration of various data centers; one of which was the BSC, which performed a commendable task being one of the earliest to advance results, executing much of the analysis and storing over 500TB of data.
|
More information on the tools and their capabilities is available from The PCAWG Data Portals and Visualizations Page (http://docs.icgc.org/pcawg).
Article: A user guide to the online resources for data exploration, visualization, and discovery for the Pan-Cancer Analysis of Whole Genomes project (PCAWG).