IBM-BSC News
The IBM-BSC team has presented libPRISM, an infrastructure that transparently manages the configuration of architectural knobs (SMT, DVFS and data prefetcher) according to the demands of parallel workloads, adapting the system to the evolving demands of hardware resources. LibPRISM can also be used to investigate possible behavior changes due to hardware knob reconfigurations and quickly prototype an algorithm to take advantage of the observed behavior. The source code of libPRISM is freely available under a BSD 3-Clause License at https://github.com/criort/libPRISM.
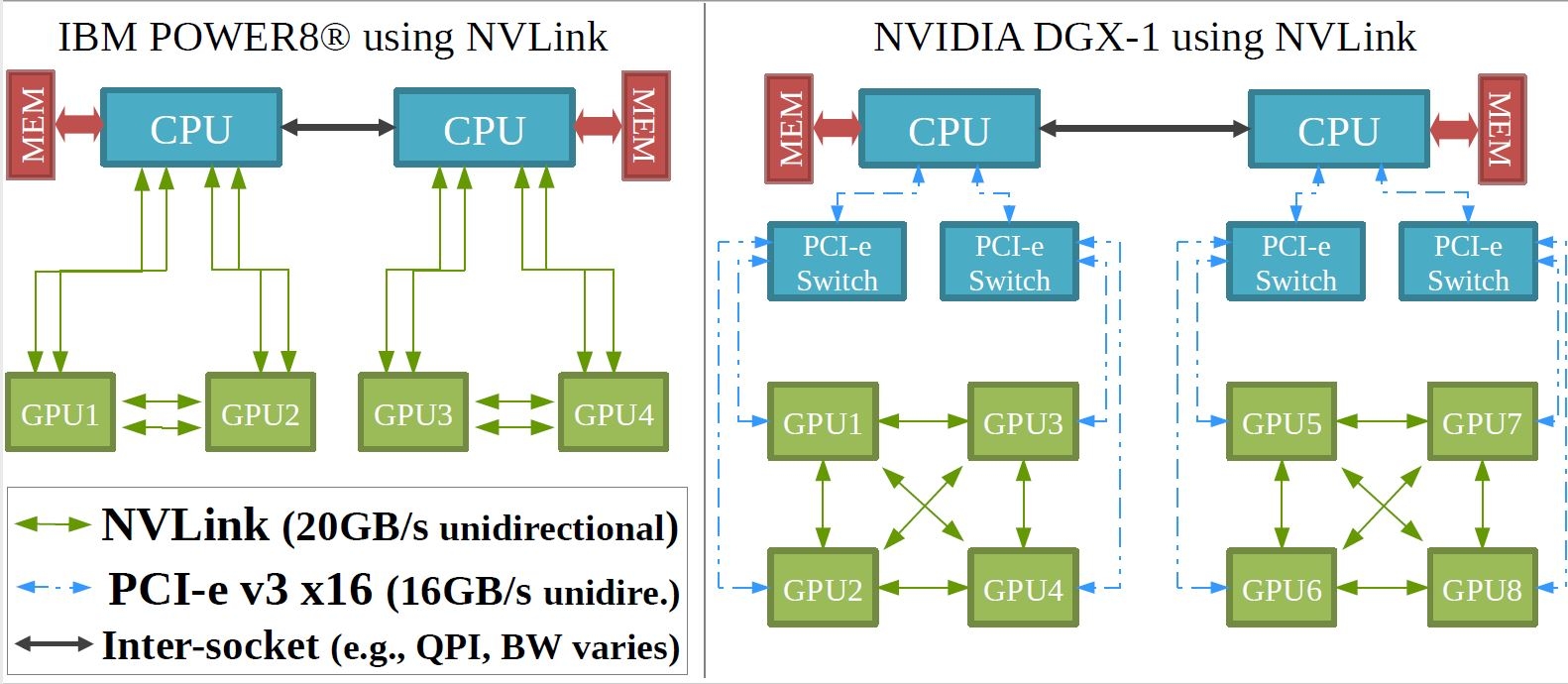
In this joint publication, researchers present a novel approach to leverage GPU disaggregation in Cloud environments. The joint team of BSC and IBM researchers presents a new topology-aware workload placement strategy to schedule deep learning jobs on multi-GPU systems. The placement strategy is evaluated with a prototype on a Power8 machine with Tesla P100 cards, showing speedups of up to ≈1.30x compared to state-of-the-art strategies; the proposed algorithm achieves this result by allocating GPUs that satisfy workload requirements while preventing interference. Additionally, a large-scale simulation shows that the proposed strategy provides higher resource utilization and performance in cloud systems. This contribution is especially relevant for cloud providers offering accelerators through XaaS solutions and running Kubernetes-based infrastructures.
As a leader in information technologies, IBM is promoting a concerted effort with researchers at the BSC Life Sciences and Computer Sciences departments towards the creation of a platform for biological ontologies interoperability. Along with large-scale parallelism and scalability, this framework harnesses machine learning to allow knowledge interconnection and discovery. The inference of a fine mapping between disease-related phenotypic features and distinct molecular processes meets applications that range all biomedical fields, from functional genomics to systems medicine, as well as possible extensions to IBM cognitive services.
IBM and BSC researchers received the best paper award for "Adaptive Pattern Matching with Reinforcement Learning for Dynamic Graphs" at HiPC 2018 conference. In this joint publication, researchers presented an approximate incremental graph pattern matching (IGPM) algorithm for dynamic graphs and its performance optimization system (PEM) using reinforcement learning and graph clustering algorithms. The paper demonstrated great performance speedup by achieving 10.1 times faster than an existing graph pattern matching algorithm using various graph structure data. This research has an impact on IBM products/business in that it provides the potential to more effective real-time graph analytics for large-scale time-evolving networks that is previously unattainable due to the computation complexities. Moreover, the proposed algorithms can be applied to various real-world graph data, such as other social network analytics and finding suspicious financial transaction detections. The paper is available at https://arxiv.org/abs/1812.10321
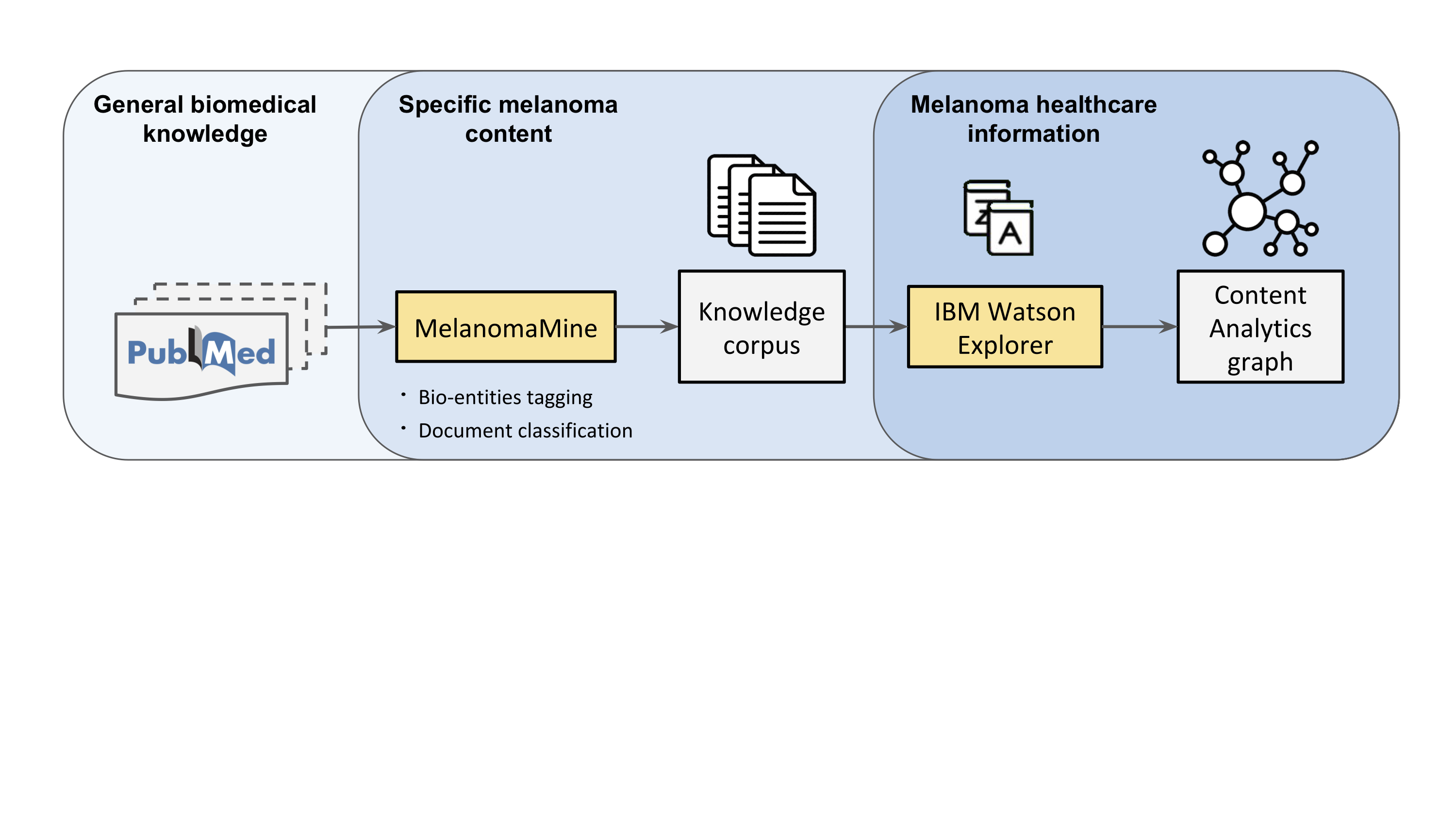
Researchers at the BSC Life Sciences Department have collaborated with IBM Research Zurich in a joint effort to put text analytics at the forefront of biomedical research in melanoma, the deadliest form of skin cancer. Thanks to MelanomaMine, a text mining application developed at BSC that uses machine learning to extract melanoma-specific information, operations of the IBM Watson cognitive computing system were enabled to provide insights into melanoma healthcare. In particular, by coupling MelanomaMine and IBM Watson, the researchers identified relevant gene-to-phenotype associations that could be used to inform melanoma risk assessment.
BSC, in collaboration with IBM, has implemented a set of linear iterative solvers leveraging tasking constructs to improve performance. The improvements achieved by BSC provide parallel performance improvements up to 42.9% and average improvements of 11.8% with respect to the best state-of-the-art techniques while keeping similar numerical stability properties. You can download the code from: https://repo.hca.bsc.es/projects/parallel-apps/repository
The CTE POWER9 cluster is one of emerging technology clusters that are part of current Marenostrum 4. This cluster is composed of 54 nodes, each one equipped with 2 Power9 processors (20 cores per processor) and 4 NVIDIA V100 GPUs. This machine contributes with 1.5 PFlops to the total BSC's computation power. Currently the machine has more than 100 applications and libraries ported to the system, making the system ready to be used by the scientific community; the main users are coming from Life sciences groups (using codes like AMBER, NAMD, GROMACS or LAMMPS), Earth sciences teams (mainly using the system to run data post-processing codes adapted to the POWER9 architecture), and engineering codes (like BSC's ALYA) which are optimized to use the POWER9 architecture and attached GPU accelerators.
From beginning 2019, the RES (Spanish supercomputing network) provides 30% of the MareNostrum4 CTE POWER9 machine to scientific researhers; this means around 1 Million core-hours per each period (with a duration of 4 months). During this first year the interest in using the machine has stedeadily grown; in fact, the demand has exceeded 2 times the resources available, requesting more than 6.3 Million hours vs the 3 Million hours available. The main scientific areas that are currently asking resources in the CTE POWER9 + NVIDIA V100 cluster are Medicine and Life sciences, followed very close by Physics and Earth sciences.